| A combinatorial set of molecules is formally generated by combinatorially placing substituents at selected positions of a core structure (invariant part). Combinatorial representations of molecule sets are common in the chemical patent literature, combinatorial chemistry and in context with QSAR studies. | ||
| Here we are choosing the Markush structure example of Barnard and Dawns that also was selected by Maclean and Martin [1] to discuss combinatorial library representations. This example demonstrates four forms of variation in generic structures: | ||
|
||
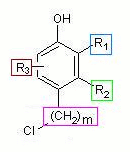
The following is a corresponding CurlySMILES notation: Oc1c{+Rcc=C{-},CC{-}}c{+R}c(C{-}{+nn=1-3}Cl)cc1{+Yc=N{-};p=6}
The core structure is phenol, represented by the SMILES code in black. The four variations are represented as OPAM annotations. The substituent variation in ortho position to the hydroxy group is included via annotation marker +R indicating alkyl group
substitution. The two substituents C{-}
(methyl) and CC{-} (ethyl) are
given as a comma-separated pair via key
cc.
The homology variation in meta
position to the hydroxy group is included by solely using
annotation marker +R, representing
a potentially unlimited class of alkyl groups.
The position variation of the
amino group at the other ortho and para positions
is encoded as +Y
(for “any group”) annotation, which is anchored
at atom position 7 of the SMILES notation (ortho position).
The second position possibility for the amino group is
atom position 6, specified via key
p.
The frequency variation, which defines a
range for the occurrence of a structural repeat unit (SRU), is
encoded by using the CurlySMILES format for an SRU
(in this case a methylene group).
|
||
| A typical combinatorial library is formally generated by substituent and/or position variation. Frequency variations are less likely to occur in synthesized libraries. But any type of variation can be useful to define a virtual library. The given example demonstates that CurlySMILES supports efficient serialization of diverse structurally defined libraries for network transport, querying and the study of interlibrary relationships. | ||
Tweet
_
__
__
 __
__
__ ___ 
|
References
| [1] |
D. Maclean and E. J. Martin:
On the Representation of Combinatorial Libraries.
J. Comb. Chem.
2004,
6 (1)
1-11. doi: 10.1021/cc0340325. |
| [2] | A. Drefahl: CurlySMILES: a chemical language to customize and annotate encodings of molecular and nanodevice structures. J. Cheminf. 2011, 3:1; doi: 10.1186/1758-2946-3-1. |

Custom Search
|